Genetic Choreography of the Developing Human Embryo
Years ago, when I was teaching at a state university, I had the privilege to show real human embryos and fetuses to my genetics classes. An obstetrician back in the 1950s had saved them after patients had miscarried, with permission I was told, and donated the collection to the biology department.
My students were astonished at the forms that floated in size order in their test tubes and flasks, culminating in an 8-month fetus in a giant mayonnaise jar. I handled them with great care and respect.
Once when I was wheeling the collection across campus in a shopping cart to get to class, students accosted me, assuming I was a right-to-lifer en route to an event. No, I explained, they were for a biology class. And one day I brought my 4-year-old to campus and she saw the 8-month fetus and burst into tears – it looked too much like her baby sister.
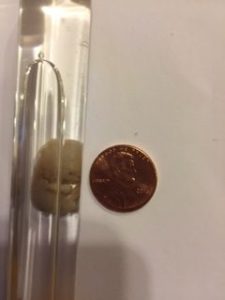
Years later, I found one of the precious test tubes wrapped in paper towels in a box of exam papers. I didn’t take it intentionally. I put it in a special place and unwrapped it for this post. The embryo had been collected on day 44, and it appears to the right next to a penny for perspective.
VIEWING ORGANOGENESIS
Human embryos and fetuses present powerful images. For my students, seeing the real thing was unforgettable in a way that illustrations, photos, and films can’t possibly be. Today, billboards near my home depict happy babies with tag lines announcing at which week a heartbeat, fingerprints, and smiles emerged, presumably to lay a guilt trip on women who must choose to end pregnancies. The messages make me yearn for a can of spray paint.
Embryologists (aka developmental biologists) have traditionally used a classifying system of 23 prenatal “Carnegie stages” based on physical characteristics visible by certain dates. A day 32 human embryo, for example, is 4 to 6 millimeters long and has buds for legs, pits for ears, thickenings in the outer layer that will become lenses, and 30 body segments (somites) that will develop into specialized body parts. By day 56, the embryo is 27 to 32 mm long, and the head now comprises half of the body. It has a chin, extended limbs fringed with fingers and toes, and the inklings of genitalia.
The period of the embryo lasts from fertilization until day 58, the end of week 8. The two-week mark is when the three primordial tissue layers (ectoderm, endoderm, and mesoderm) emerge and then bend to form the classic gastrula, from which organs unfurl and elaborate according to a precise genetic program.
Starting around week 3, the rudiments of organs begin to form, and a week later this period of organogenesis accelerates. The embryo/fetus transition by the end of the eighth week (counting from conception, not the confusing “last menstrual period” shortcut of obstetricians) is when all the precursor structures are present. Beware posters at Planned Parenthood protests and in the media that conflate embryo and fetus – they are biologically quite distinct stages of prenatal development. I’ve even seen printed materials at obstetrician offices referring to an embryo or fetus as a baby.

Carnegie staging is based on what’s visible. But the genome controls the launching of the observable anatomical changes that drive the journey from embryo to fetus. A stunning recent paper in the journal eLife, from Dave T. Gerrard and Neil Hanley and colleagues from the University of Manchester and Central Manchester University Hospitals, National Health Service Foundation Trust in the UK, offers an entirely new view of the human embryo based on gene expression in and across specific organs.
The investigation differs from past research directions to probing prenatal development, such as analyzing RNA from whole embryos, observing stem cells as their daughter cells differentiate, animal models of human diseases, creating lab-nurtured human organoids, and inferring the developmental detours behind certain birth defects.
TRACKING GENE EXPRESSION AS EMBRYOS DEVELOP
The report, “An integrative transcriptomic atlas of organogenesis in human embryos,” weds the statistical tool of principal component analysis (PCA) to cell lineage pathways that extend as the initial cleavage cell divisions and tissue layering become organogenesis. PCA reduces huge data sets to a manageable number that represent trends or substructures within the many variables. The approach, which the researchers call LgPCA for “lineage guided PCA,” identifies “metagenes,” which are groups of genes that partake in the formation of specific body parts in the embryo, some of them multi-tissue.
The new work catalogs RNAs that are transcribed from the genes that are accessed as development proceeds from the end of the third week until the end of the eighth. The research was possible because of the decades-long legacy of ethically obtaining embryo tissue from women undergoing voluntary pregnancy termination in the UK. The Human Tissue Authority website describes every nuance of working with human cells, tissues, and organs. It includes bioethical concerns such as consent and dignity, displays in museums, religious and cultural ideas about donating organs, use of stem cell and other cell lines, and use of embryonic and fetal tissue from pregnancy terminations.
The researchers collected material and then dissected out 15 specific parts for analysis. The specimens included whole organs such as adrenal glands as well as distinct segments, such as the retinal pigment epithelium in the eye, the stomach unhinged from its sphincters, and limb buds. The investigators pooled parts to isolate enough RNA to analyze and identified the genome regions from which they were transcribed.
6,000+ “TRANSCRIPTIONAL CODES”

By isolating RNAs from distinct body parts, the researchers could identify the DNA sequences behind complex congenital anomalies that involve more than one tissue type, such as cleft palate and some forms of congenital heart disease.
The most unexpected finding was that about 90 percent of 6,251 novel RNA transcripts identified are not the usual protein-encoding variety at all, but are “long intergenic non-coding RNAs,” aka “LINC RNAs.” They are highly specific to body parts, where they apparently control the deployment of transcription factor proteins, which in turn steer the program of gene activation and suppression that directly oversees the changes of development.
The huge dataset yielded 11 metagenes. An example is “metagene 2,” which represents 39 genes that sculpt the embryonic liver. Because the LgPCA approach is based on cell lineages, it can reveal the genetic programs behind developmental syndromes, such as Holt-Oram syndrome that affects the limbs and heart.
Knowing these genetic controls behind organogenesis can perhaps lead to development of diagnostic tests or even identify drug targets. On the research front, knowing the gene expression patterns and signals that herald an event – a spleen forming, for example – might establish benchmarks that can be used in the creation of induced pluripotent stem cells and coaxing them towards specialization.
THE BIGGER PICTURE
I am utterly fascinated with this new lens into organogenesis. The researchers describe the impact of their work best: “the discovery of a major new programme of non-coding transcription adds a fresh layer of detail on the spatiotemporal regulation of the human genome.”

I think the identification of a role for non-protein-encoding RNAs in sculpting the organs of the embryo is on a par with that of the finding that genes are in pieces – the introns snipped out, leaving only the exons to encode protein – articulated so beautifully by Walter Gilbert back in 1978. The discovery of introns instantly dispelled the long-held view of the gene as a single, sleek, DNA code for an RNA molecule.
Like introns, LINC RNAs were also once not well understood because they do not encode protein. The wonder of genetics is that we often think that we know close to all there is to know, only to discover yet another hidden language of life.
